About the Pathway Prediction System
 Overview
Overview
The EAWAG-BBD Pathway Prediction System predicts microbial catabolic reactions using substructure searching,
a rule-base, and atom-to-atom mapping. The system is able to recognize organic functional groups found in
a compound and predict transformations based on biotransformation rules. The biotransformation rules are based on
reactions found in the EAWAG-BBD database or in the scientific literature.
The pathway prediction system can be accessed at the EAWAG-BBD Pathway Prediction
page, which can be reached from the "Pathway Prediction" link on the EAWAG-BBD home page,
or by using the following URL: http://umbbd.ethz.ch/predict/.
The PPS predicts biodegradation of user compounds. Biodegradation of some types of compounds should not be predicted. Users can
choose if they will view all or only the more likely aerobic transformations.
The PPS uses Chemaxon's MarvinSketch and MarvinView Java applets as plugins.
A user enters a compound into the system by one of two methods: Drawing the Structure and Generating SMILES
or Entering SMILES Directly.
 Demonstration
Demonstration
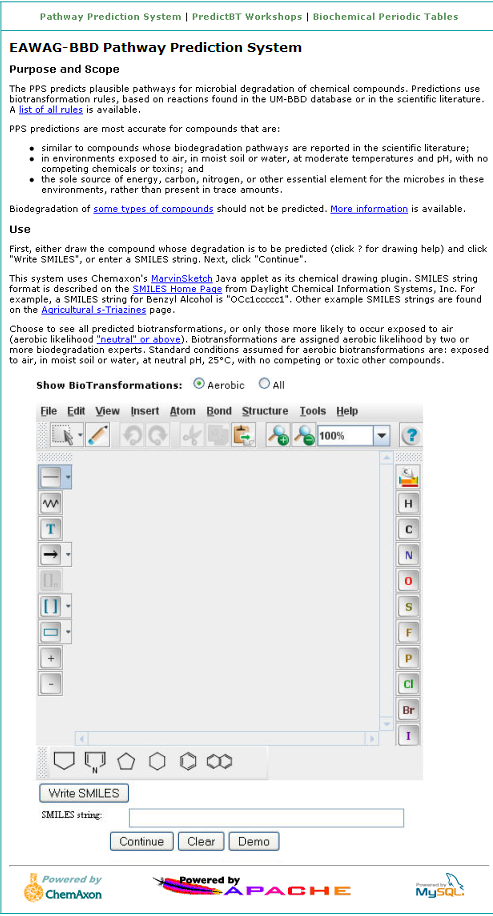
The PPS provides a demonstration of how the system works using the compound Benzyl Alcohol. To access the
demo, click on the "Demo" button located under the MarvinSketch window of the
EAWAG-BBD Pathway Prediction Page.
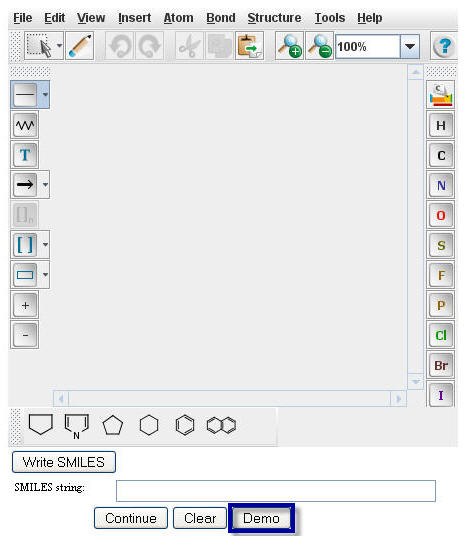
The Smiles string of Benzyl Alcohol will appear in the "Smiles string" box.
 Status Page
Status Page
A user will be automatically taken to a prediction status page after clicking on the "Demo" button as shown above, or otherwise submitting a query compound, and clicking on the "Continue" button.
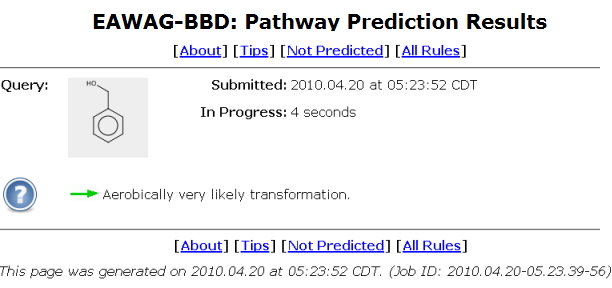

The query compound is shown in a grey square in upper left of the status page.
The query submission time and the prediction run-time are shown next to the compound square.
The prediction run-time is refreshed once a second.
A system usage tip is displayed under the compound square.
Different random tips are shown every five seconds.
A
list of all tips is available.
A unique job id is shown at the bottom of the status page.
When the initial prediction on the query compound is finished, the system will bring a user to the Pathway Prediction Results page.
 Pathway Prediction Results
Pathway Prediction Results
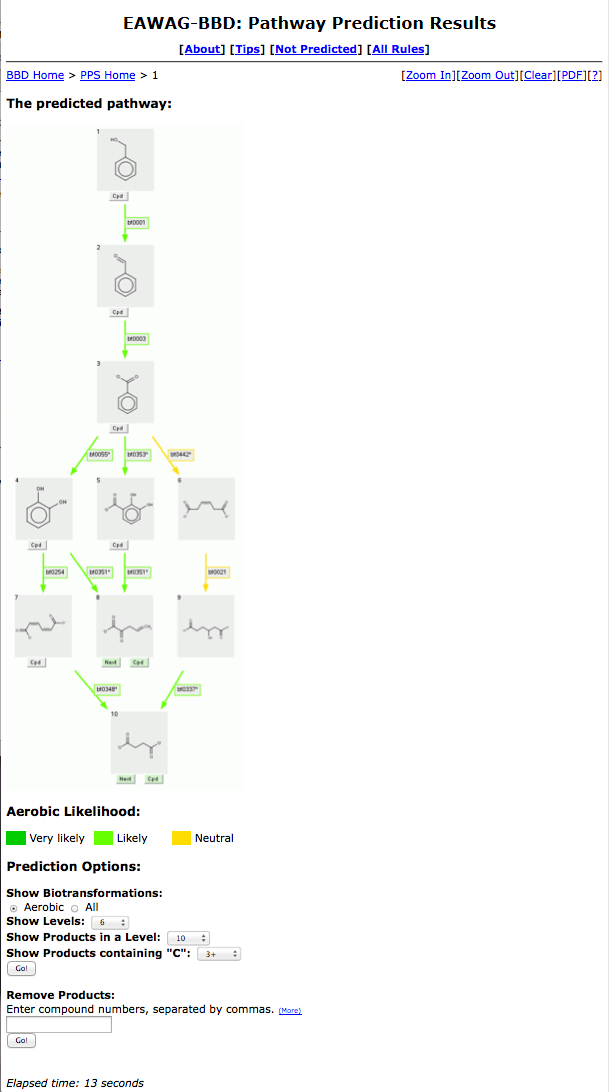
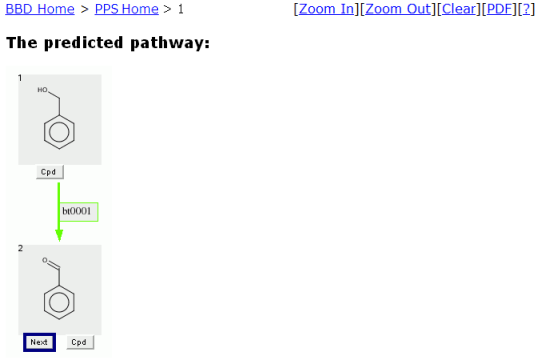
The Pathway Prediction Results page for Benzyl Alcohol is shown below using default prediction options.
Each numbered grey square contains one compound.
The default options are changeable through out the prediction process as described under "
Change Default Options" below.
From the Pathway Prediction Results page, a user is able to see the more likely aerobic transformations of Benzyl Alcohol based
on the biotransformation rules found in the EAWAG-BBD.
The Navigation Bar "BBD Home > PPS Home > 1" permits quick navigation back to previous prediction steps and the PPS and BBD home pages.
The Zoom feature to the right of the Navigation Bar allows the user to resize the prediction results as described on the Viewing and Printing page. The prediction run-time is shown in the "Elapsed time" at the bottom of the results page.
 MarvinView
MarvinView
The user can click on any grey compound square in the predicted pathway.
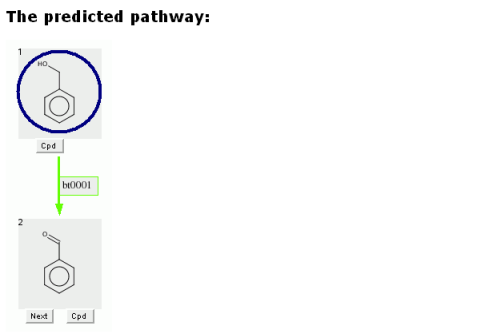
Clicking on a grey compound square will show the compound in a new window.
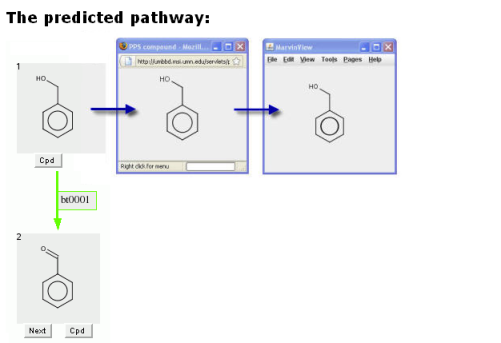
The compound can be rotated in this window. The window can also be increased in size, to better view larger compounds.
Double clicking in this new window will produce a MarvinView window.
The MarvinView window can provide much additional information on the compound.
Information on how to use it is available at
http://www.chemaxon.com/marvin/help/view/view-index.html.
 Change Default Options
Change Default Options

By default, the PPS will show "Aerobic" biotransformations,
show up to 6 levels,
display compounds containing 3 or more carbon atoms,
and stop before any level where there are more than 10 compounds.
These default options are changeable throughout the prediction process, as described below.
 Aerobic Likelihood
Aerobic Likelihood
Aerobic Likelihood is the likelihood that the reaction will occur under aerobic conditions, exposed to air, in soil (moderate moisture) or water, at
neutral pH, 25°C, with no competing or toxic other compounds.
Rules are assigned aerobic likelihood by
two or more biodegradation experts.
More information is available.
The aerobic likelihood of a transformation is indicated by a color of an arrow that starts from one substrate to one or two products.
Users are able to choose, before or after a compound undergoes transformation, if they will view all
or only the aerobic likely transformations of a compound by selecting the appropriate button.
If the aerobic setting is chosen, and no very likely, likely, or neutral transformations, but some unlikely and/or very unlikely transformations, are found, the setting is automatically changed to "All" and these transformations are shown.
 Show Levels
Show Levels
By default, the system will predict up to 6 levels in the first step.
The "All" choice is shown below for toluene, an immediate precursor of the benzyl alcohol used in the above examples, in a prediction set to 1 level.
If "Aerobic" were chosen, only the first two choices (3-methylcatechol and benzyl alcohol) would be shown.
When "All" is chosen, a third choice, succinate addition to the toluene methyl side chain, is displayed.
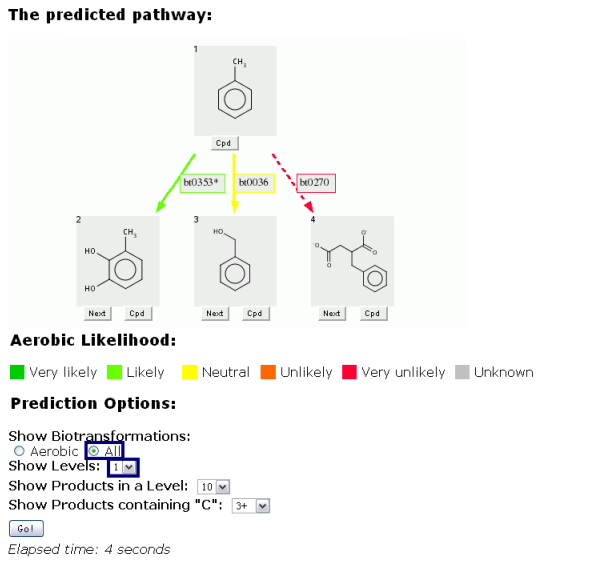
Some predictions are too complex to be displayed. When this occurs, the prediction is rerun with a decreased number of levels, and a message is displayed under the last compound row as shown below.
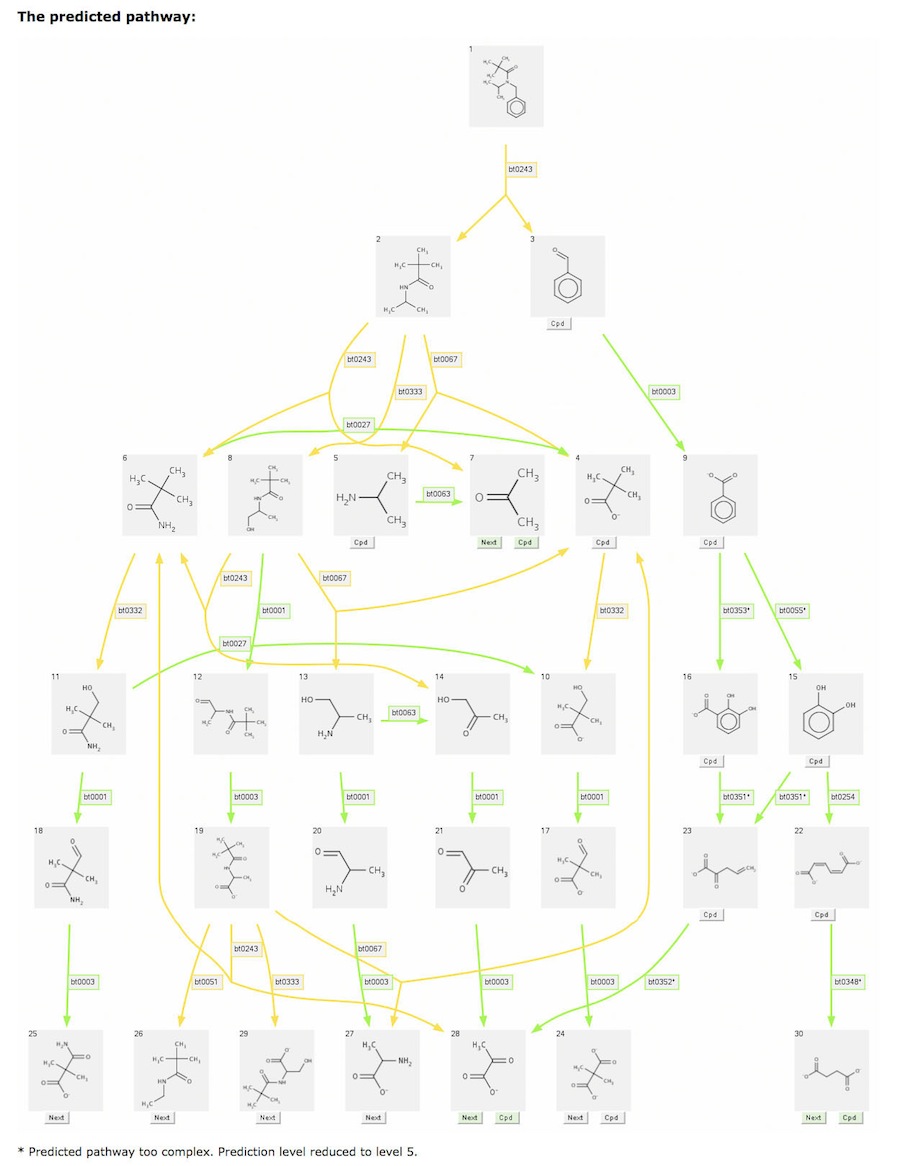
 Show Products Containing "C"
Show Products Containing "C"
The next examples show predictions for the herbicide malathion [CCOC(=O)CC(C(=O)OCC)SP(=S)(OC)OC], a compound with many organic functional groups, including two ethyl esters.
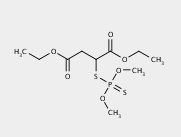
The first three levels of the prediction for malathion are shown below using default prediction options.
By default, the system will only display compounds containing 3 or more carbon atoms. A user can change this minimum number of carbon atoms to view more or fewer compounds.
In the malathion prediction shown above, the ethanols cleaved by bt0024, the ester cleavage rule, are not displayed using the default settings.
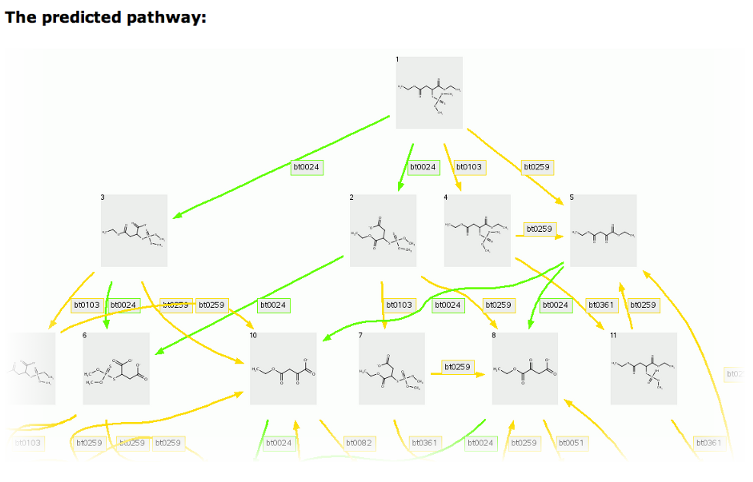
The 1-level prediction for malathion with an option to show 2 or more carbon atoms, which shows the missing ethanol, is shown below.
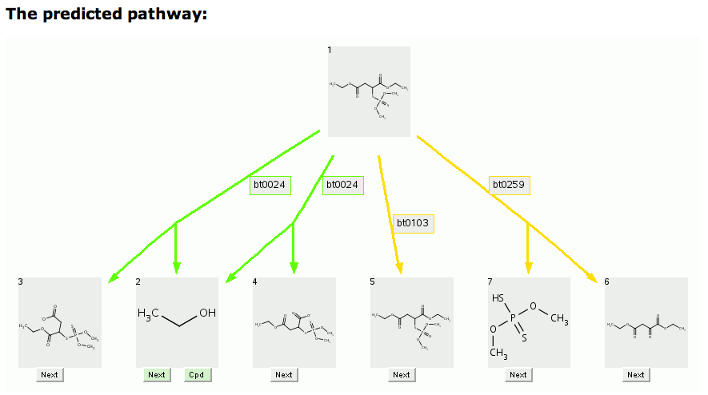
Cleavage compounds (Compounds 2 and 3, Compounds 2 and 4, Compounds 6 and 7) are indicated by two tailed arrows. Duplicate compounds are removed and only one (Compound 2) is left, pointed to by multiple arrows.
An exception to "Show compounds containing above a threshold number of carbon atoms" occurs when the parent compound is one carbon or less above the threshold.
In those cases, compounds with one or more carbons are shown.
 Show Products on a Level
Show Products on a Level
By default, the system will display up to 10 compounds on a level and stop before any level where the number of compounds exceeds this threshold.
If a prediction is stoped due to the threshold, a message will be shown under the last compound level, as shown below.
All compounds in the very first level of any prediction step will be shown, even if they exceed the threshold.
A user can decrease or increase the threshold value.
Shown below is the prediction for malathion, limited to no more than 5 compounds.
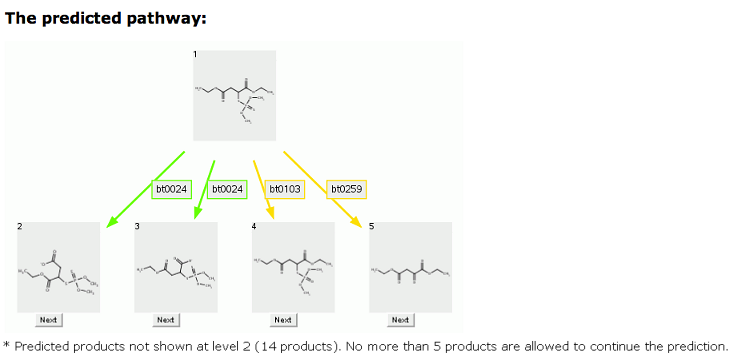
 Rule Button
Rule Button
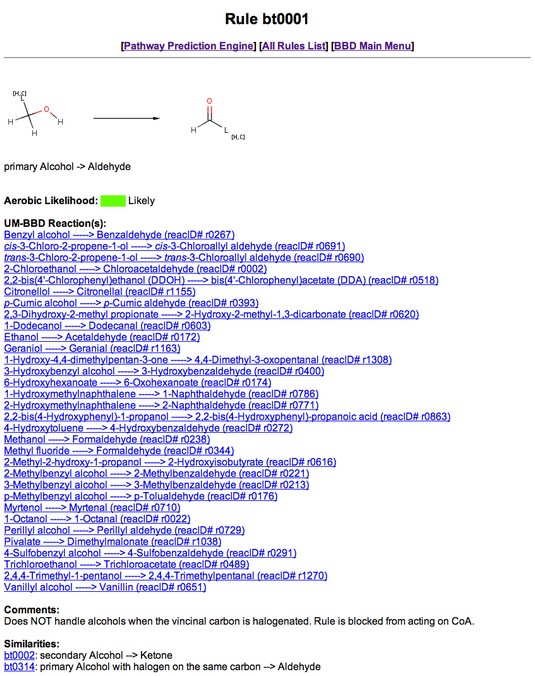
The demo provides the user with the ability to click up to three buttons for each transformation.
One of these, always shown, is the "Rule" button.
The "Rule" button is shown as the label of an arrow which represents a predicted transformation.

Clicking the "Rule" button will open a new browser window containing the rule page of the biotransformation rule associated
with the specific reaction. The Biotransformation rule page provides a user with links to the PPS, a list of
All BTrules, and the EAWAG-BBD Main Menu. It also provides one or more descriptions of the type(s) of reactions
handled by the rule, the likelihood the reaction will occur under aerobic conditions, a list of EAWAG-BBD reactions
that exemplify the rule, staff comments, similar biotransformation rules (if any), and a way for the user to provide
comments. A
list of all rules the system uses is available. More information about BTrules is available in
About the BBD.
 Cpd Button
Cpd Button
Another button is the "Cpd" button, located in the bottom of an individual compound square.
The "Cpd" button does not appear on every transformation (only on compounds found in the EAWAG-BBD).

Clicking the "Cpd" button will open a new browser window containing the compound page of a biotransformation. The compound page
provides a user with a graphic image of the compound, its chemical formula, molecular weight, SMILES string,
and links to reaction pages in the EAWAG-BBD where the compound is either the substrate or product. More information about compound pages is available in
About the BBD.

 Next Button
Next Button
Another button is the "Next" button, also located in the bottom of an individual compound square,
to the left of the "Cpd" button, if one is present.
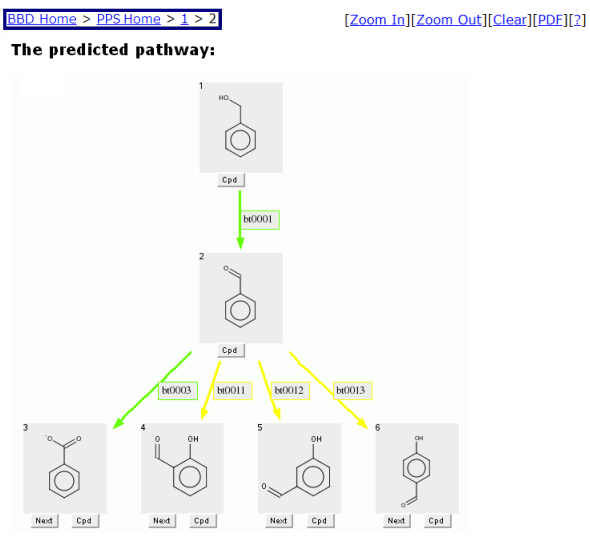
To continue the pathway, select the "Next" button on another transformation. The Navigation Bar will then expand to "BBD Home > PPS Home > 1 > 2" for easy navigation to previous PPS prediction pages or BBD or PPS Home pages.
The biotransformed compound selected will appear as an intermediate compound in the new predicted pathway
starting from the original compound.
The colored arrow indicates the aerobic likelihood of the biotransformation that connects two compounds.
 Green Next or Cpd Button
Green Next or Cpd Button
After several cycles of the prediction process, you might see a green-shaded version of the "Next" button.
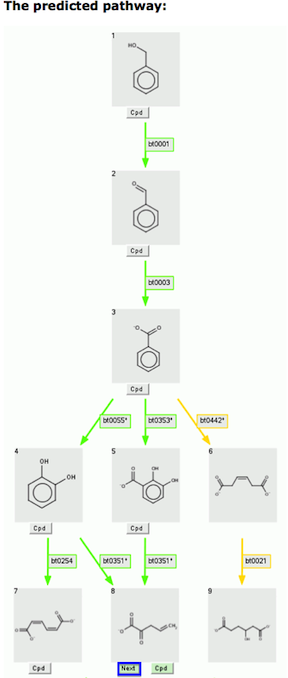
This indicates that this compound is readily degraded. If you click on a "Green Next" button, your predicted pathway is completed by ending at the chosen compound.
At the same time, all branches ending at other compounds are removed.
In the example below, compound 6 and 8, not on the
path between compound 1 and compound 9, the chosen compound, are removed.
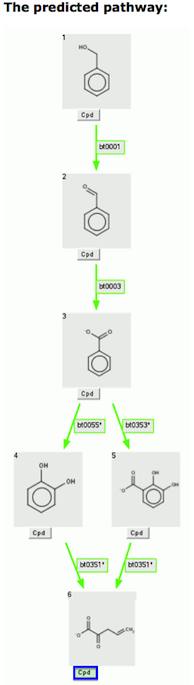
The "Green Cpd" button indicates that this compound is found in a KEGG pathway. Clicking the "Green Cpd" button brings a user to a KEGG pathway web page. Alternatively, a user may choose to continue prediction by clicking on a non-green "Next" button, if one is present.
 Remove Products
Remove Products
As shown above, when the Next button is clicked, the new pathway contains that compound and the pathway branch or branches that produced it.
That is, other predicted compound squares and rule arrows are removed.
The "Remove Products:" option at the bottom of a prediction, allows more control over compound removal.

Compounds are removed by entering their numbers (the small number shown in the upper left of each grey compound square), separated by commas, in the "Remove Products:" box, and then clicking on the "Go!" button below the box.
The starting compound (compound 1) cannot be removed.
The
(More) link will open this section of the EAWAG-PPS help webpage in a separate window.
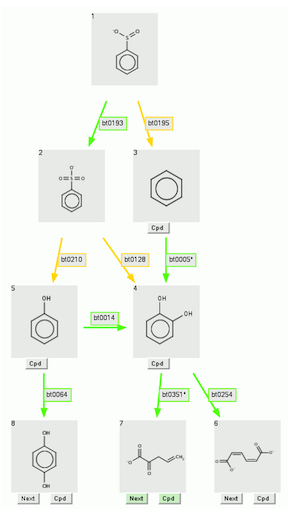
For example, above is the 3-level prediction for benzenesulfinate.
If it is desired to remove the compound benzene (product 3) in level 1, and phenol in level 2 (product 5), these compounds, and all compound boxes and rule arrows following them, are removed
by entering "3,5" (or "5,3") in the "Remove Products:" box, and clicking the "Go!" button.

The partial prediction is shown below:
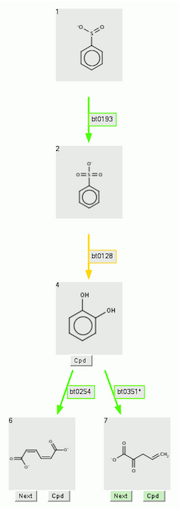
To return to the original prediction, click on the "Show Original" button, to the right of the "Go!" button after a partial prediction has been made.

Once back at the original, you can return to the partial prediction by clicking on the "Show Partial" button, in the same position as the "Show Original" button.
Compounds in a partial prediction retain the same numbers as in the original prediction.
Since some compounds were removed, there are gaps in the number sequence.
Removing compounds can be done in multiple steps, without returning to the original prediction.
In the above example, you can get to the same result by first choosing to remove compound 3 (or 5),
and, after reviewing the result, choosing to remove compound 5 (or 3).
If you return to the original prediction, you can remove different compounds to create a new partial prediction.
If a previous partial prediction exists, the new one overwrites it.
Zoom In, Zoom Out, and PDF buttons all work in a partial prediction the same as in the original.
Clicking the Clear button in a partial prediction returns to the original.
Compound, Rule and Next buttons work in a partial prediction as in the original.
If the Next button is clicked in a partial prediction, the Next step in the prediction will be that of the original, not the partial, prediction.
 Variable Aerobic Likelihood
Variable Aerobic Likelihood
Some rules handle a wide range of compounds, and a single aerobic likelihood is not appropriate for all of them.
Thus structure-based variable aerobic likelihood has been added to the PPS.
For aerobically likely aromatic ring monooxygenation, ring dioxygenation, ring cleavage, and ring decarboxylation rules, when the substrate contains one or more halogens, the aerobic likelihood for the transformation is changed to "neutral".
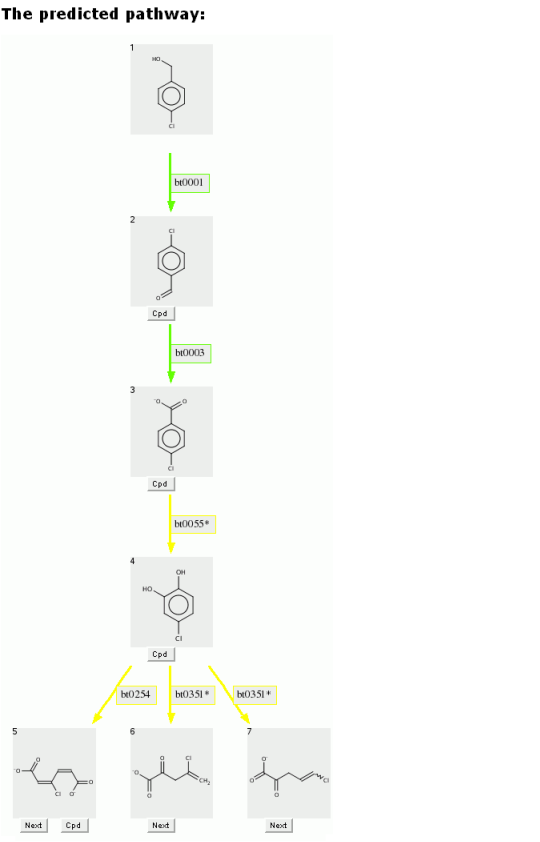
Compare the previous predicted pathway for benzyl alcohol, with the pathway below for p-chlorobenzyl alcohol.
The aerobic likelihood for the first three transformations are likely, as above.
However when the ring is oxygenated and cleaved, the aerobic likelihoods are neutral for the chlorinated compound, rather than the likely value found for the non-halogenated compound.
A second use of variable aerobic likelihood is for nitro groups. Aerobically likely or neutral rules for oxidation of aliphatic carbons have been changed to unlikely when the chain contains one or more nitro groups.
 Super Rules
Super Rules
Super rules include two or more contiguous biotransformation rules that form a small pathway of their own. They are indicated with asterisk. For example, bt0055 and bt0351 are super rules shown on the Figure in "
Variable Aerobic Likelihood" section above.
 Drawing Structures
Drawing Structures
The PPS uses Chemaxon's MarvinSketch for drawing compounds. To draw Benzyl Alchol in the MarvinSketch
window, select the "Benzene Ring" from the lower tool bar menu. Drag the structure to the desired location in
the window and click to paste. Next select the "single bond" button from the left tool bar (for more options e.g. double bond, single
bond up, etc., click and hold the "single bond" button). With the "single bond" selected, click a corner of
the bezene ring, drag the bond in the desired direction and release to paste. With the "single bond" still
selected, click the methyl group, drag in the desired direction, and release to create a two carbon chain.
To make the compound an alcohol, select the atomic symbol for oxygen, "O", from the right menu bar. Click on the
methyl group to replace it with a hydroxide. For further information, a tutorial for using Marvin Sketch is
available at
http://www.chemaxon.com/products/marvin/marvinsketch/.
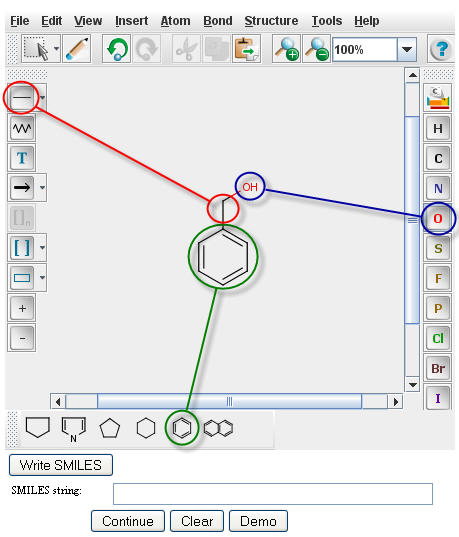
After creating an image image in the Marvin Sketch window, a user must generate the SMILES string for that
specific compound. To generate the SMILES string, click the "Write SMILES" button located beneath the
MarvinSketch window.
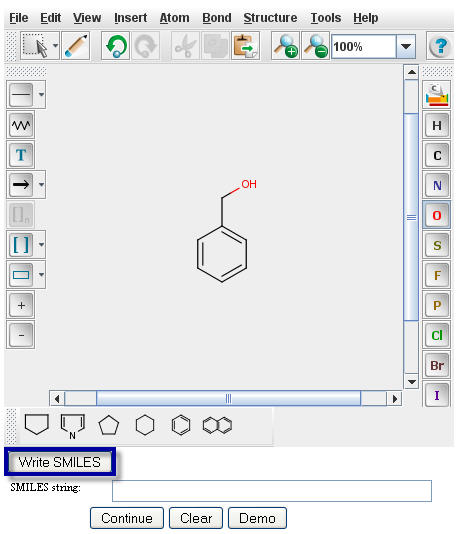
The SMILES string will appear in the window directly underneath your structure.
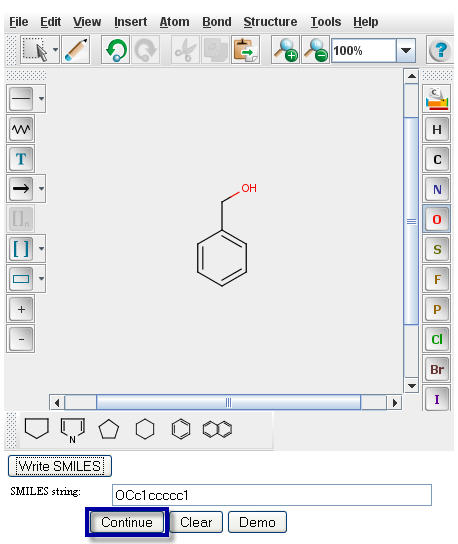
Click on the "Continue" button to proceed to the
Pathway Prediction Results page for Benzyl Alcohol.
 Entering SMILES Strings
Entering SMILES Strings
If your brower cannot run Java applets, or you have the SMILES string for your compound, you can instead
enter a SMILES string. A SMILES string is an alphabetical representation of a compound.
The SMILES string format is described on the
SMILES Home Page from Daylight
Chemical Information Systems, Inc.
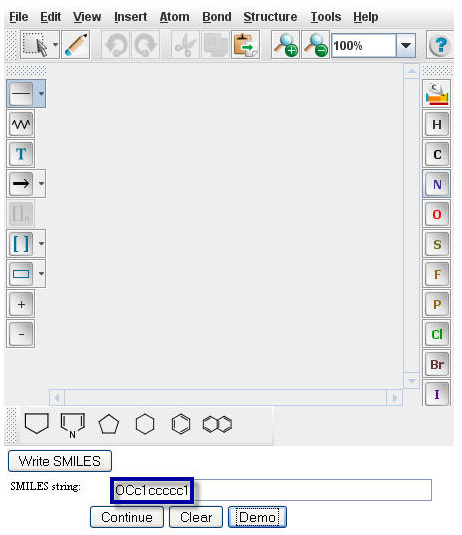
Type or paste the SMILES string of Benzyl Alcohol, OCc1ccccc1, into the "Smiles string" window.

Click on the "Continue" button to proceed to the
Pathway Prediction Results page for Benzyl Alcohol.
 For Further Information or To Make Comments
For Further Information or To Make Comments
To receive periodic progress reports,
join our email list.
Comments related to the database are appreciated;
contact us.
